Example scripts¶
Showing 3D Directional Sensitivity¶
The 3d_dtcwt_directionality.py script in the examples directory shows how one may demonstrate the directional sensitivity of the 3D DT-CWT complex subband coefficients. It computes empirically the maximally sensitive directions for each subband and plots them in an interactive figure using matplotlib. A screenshot is reproduced below:
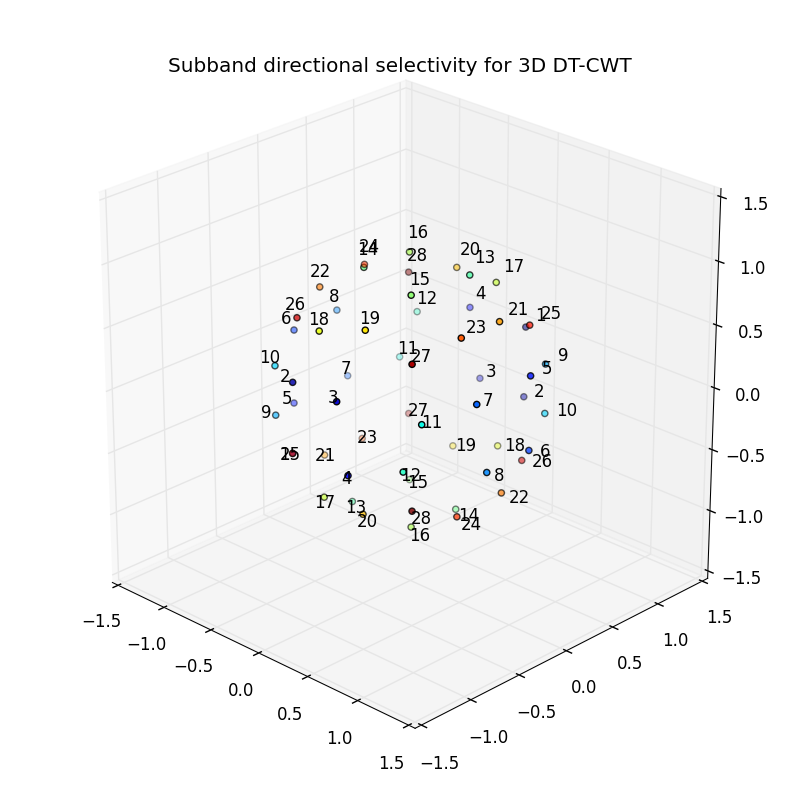
The source for the script is shown below:
#!/bin/python
"""
An example of the directional selectivity of 3D DT-CWT coefficients.
This example creates a 3D array holding an image of a sphere and performs the
3D DT-CWT transform on it. The locations of maxima (and their images about the
mid-point of the image) are determined for each complex coefficient at level 2.
These maxima points are then shown on a single plot to demonstrate the
directions in which the 3D DT-CWT transform is selective.
"""
# Import the libraries we need
from matplotlib.pyplot import *
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
from dtcwt import dtwavexfm3, dtwaveifm3, biort, qshift
# Specify details about sphere and grid size
GRID_SIZE = 128
SPHERE_RAD = 0.33 * GRID_SIZE
# Compute an image of the sphere
grid = np.arange(-(GRID_SIZE>>1), GRID_SIZE>>1)
X, Y, Z = np.meshgrid(grid, grid, grid)
r = np.sqrt(X*X + Y*Y + Z*Z)
sphere = 0.5 + np.clip(SPHERE_RAD-r, -0.5, 0.5)
# Specify number of levels and wavelet family to use
nlevels = 2
b = biort('near_sym_a')
q = qshift('qshift_a')
# Form the DT-CWT of the sphere
Yl, Yh = dtwavexfm3(sphere, nlevels, b, q)
# Plot maxima
figure(figsize=(8,8))
ax = gcf().add_subplot(1,1,1, projection='3d')
ax.set_aspect('equal')
locs = []
scale = 1.1
for idx in xrange(Yh[-1].shape[3]):
Z = Yh[-1][:,:,:,idx]
C = np.abs(Z)
max_loc = np.asarray(np.unravel_index(np.argmax(C), C.shape)) - np.asarray(C.shape)*0.5
max_loc /= np.sqrt(np.sum(max_loc * max_loc))
locs.append(max_loc)
ax.text(max_loc[0] * scale, max_loc[1] * scale, max_loc[2] * scale, str(idx+1))
ax.text(-max_loc[0] * scale, -max_loc[1] * scale, -max_loc[2] * scale, str(idx+1))
locs = np.asarray(locs)
ax.scatter(locs[:,0], locs[:,1], locs[:,2], c=np.arange(locs.shape[0]))
ax.scatter(-locs[:,0], -locs[:,1], -locs[:,2], c=np.arange(locs.shape[0]))
w = scale * 1.2
ax.auto_scale_xyz([-w, w], [-w, w], [-w, w])
legend()
title('Subband directional selectivity for 3D DT-CWT')
tight_layout()
show()
# vim:sw=4:sts=4:et